- Blog
- About
- Contact
- Jaan tere naam hd video songs download
- Tere naam movie salman khan
- Paragon ntfs western digital
- Free serial number sapphire plugins
- Video viewer apps
- How to save fabriwin nc format
- Autodesk inventor 2015 crack
- Dragon ball z wallpapers 1600x900
- Skyfonts bitstream font navigator
- Adobe premier elements 11 review
- Alaina carter modelmayhem
- Wii emulator classic systems
- How do i control link order in ncplot
- Pca column manual v6-00 pdf
- Tmobile unlock server unavailabe
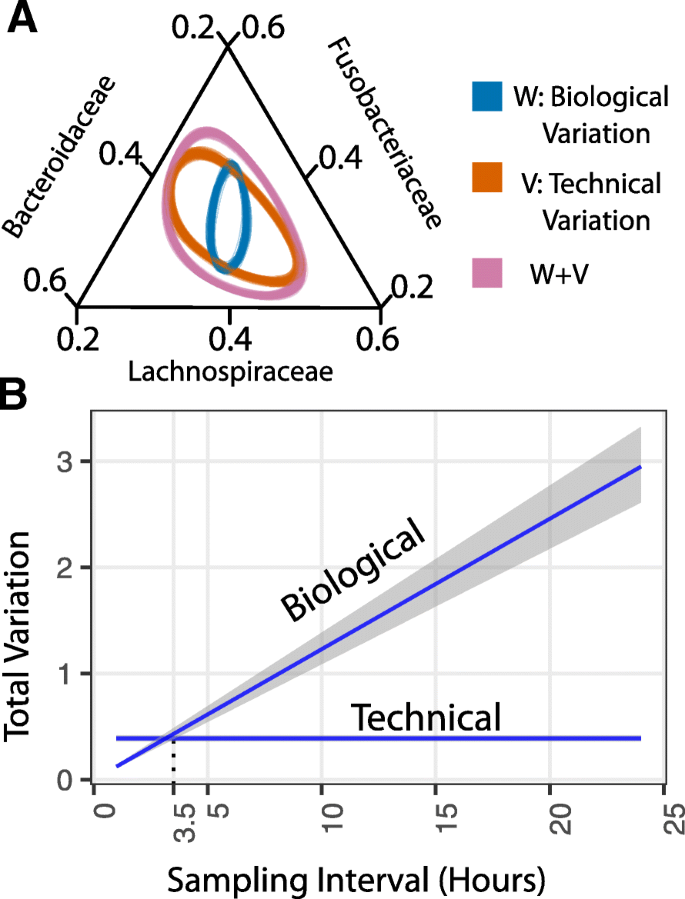
Pmatrix <- scale ( gisette_nzv ) princ <- prcomp ( pmatrix ) # change nComp to try different numbers of component variables
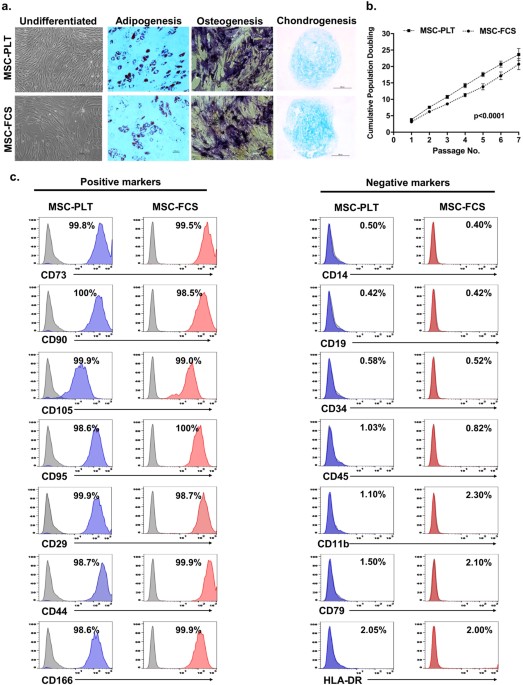
Gisette_nzv 0.1 ,])) ] print ( paste ( 'Column count after cutoff:', ncol ( gisette_nzv ))) # Run model on original data setĭfEvaluateOrig <- cbind ( as.ame ( sapply ( gisette_nzv, as.numeric )), cluster = g_labels $ V 1 ) Evaluate_GBM_AUC ( dfEvaluateOrig, CV = 5, trees = 10, depth = 2, shrink = 1 ) # Run prcomp on the data set Require ( ROCR ) require ( caret ) require ( ggplot2 ) Evaluate_GBM_AUC 0.1 ,]) # remove zero & near-zero variance from original data set NComp <- 5 nComp <- 10 nComp <- 90 nComp <- 20 nComp <- 50 nComp <- 100 # change nComp to try different numbers of component variables (10 works great)ĭfComponents <- predict ( princ, newdata = pmatrix ) dfEvaluate <- cbind ( as.ame ( dfComponents ), cluster = g_labels $ V 1 ) EvaluateAUC ( dfEvaluate )
#PCA COLUMN MANUAL V6.00 PDF FULL#
Ggplot ( dfEvaluate, aes ( x = PC1, y = PC2, colour = as.factor ( g_labels $ V 1+1 ))) + geom_point ( aes ( shape = as.factor ( g_labels $ V 1 ))) + scale_colour_hue () # full - 0.965910574495451 Pmatrix <- scale ( gisette_nzv ) princ <- prcomp ( pmatrix ) # plot the first two components Gisette_nzv 0.1 ,])) ] print ( paste ( 'Column count after cutoff:', ncol ( gisette_nzv ))) #ĭfEvaluate <- cbind ( as.ame ( sapply ( gisette_nzv, as.numeric )), cluster = g_labels $ V 1 ) EvaluateAUC ( dfEvaluate ) # Require ( ROCR ) require ( caret ) require ( ggplot2 ) EvaluateAUC 0.1 ,]) # remove zero & near-zero variance from original data set I Next nd another linear function of x, 0 2x, uncorrelated with 0 1x maximum variance.
